Trajectory SP1428
Force field:
martini_v3
Simulation length (ns): 5000
Electric field (kJ mol-1 nm-1 e-1): 0
Temperature (K): 300 (v-rescale)
Pressure (bar): 1 (Parrinello-rahman semiisotropic)
Number of particles: 19211
Time step (fs) : 25
Software: GROMACS 2021.5
Simulation length (ns): 5000
Electric field (kJ mol-1 nm-1 e-1): 0
Temperature (K): 300 (v-rescale)
Pressure (bar): 1 (Parrinello-rahman semiisotropic)
Number of particles: 19211
Time step (fs) : 25
Software: GROMACS 2021.5
Supercomputer:
Finisterrae III CESGA
Peptides: P503 NC02460
Lipids: POPC
Heteromolecules:
Ions: CL
Water model: W
Peptides: P503 NC02460
Lipids: POPC
Heteromolecules:
Ions: CL
Water model: W
Sequence :
KYKKALKKLAKLL
Total charge (e): +6
Number of residues: 13
By amino acid: Basic: 6 Acidic: 0 Hydrophobic: 6 Polar: 1 Electrostatic Dipolar Moment (e nm): 4.76
Longitudinal (e nm): 4.3 Transversal (e nm): 2.04 Hydrophobic Dipolar Moment (nm): 4.2
Longitudinal (nm): 4 Transversal (nm): 1.28 Secondary structure: Helix
Activity:
Download Files
ITP file. JSON file. PDB file.
Click on any component to highlight it from the plot.
Upper leaflet
Lower leaflet
Lipids
Membrane model for: POPC (Healthy mammal)
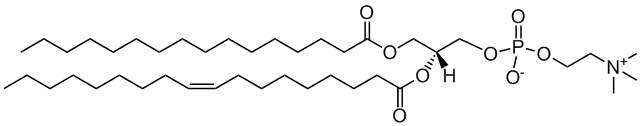
POPC
2-oleoyl-sn-glycero-3-phosphocholine
Total charge (e): 0
See POPC lipid
Download ITP File. Download PDB File.
Last snapshot
Total contacts per residue
Molecular Dynamics based descriptors
Average and standard deviation,
calculated using the autocorrelation function (for time
series)
or the width of the distribution, for the last microsecond
of
the trajectory
Area per lipid
Membrane (nm2): 0.64830300 ± 0.00121209
Upper leaflet (nm2): 0.64830300 ± 0.00121209
Lower leaflet (nm2): 0.64830300 ± 0.00121209
Average Z coordinate
Peptide (nm): 6.80835 ± 1.91528
First Residue (nm): 6.82614 ± 1.91968
Last Residue (nm): 6.79030 ± 1.90706
Membrane (nm): 6.8389100 ± 0.0123071
Upper leaflet Head Group (nm): 8.7817800 ± 0.0148239
Lower leaflet Head Group (nm): 4.89650000 ± 0.00991474
Bilayer Thickness (nm): 3.885290 ± 0.017834
Peptide insertion (nm): -1.91186 ± 1.91531
Contacts
Peptide - Water: 70.017500 ± 0.765247
Peptide - Head groups: 0.247500 ± 0.200858
Peptide - Tail groups: 0.0125000 ± 0.0271352
Tilt (°): 91.66250 ± 6.17481
Membrane (nm2): 0.64830300 ± 0.00121209
Upper leaflet (nm2): 0.64830300 ± 0.00121209
Lower leaflet (nm2): 0.64830300 ± 0.00121209
Average Z coordinate
Peptide (nm): 6.80835 ± 1.91528
First Residue (nm): 6.82614 ± 1.91968
Last Residue (nm): 6.79030 ± 1.90706
Membrane (nm): 6.8389100 ± 0.0123071
Upper leaflet Head Group (nm): 8.7817800 ± 0.0148239
Lower leaflet Head Group (nm): 4.89650000 ± 0.00991474
Bilayer Thickness (nm): 3.885290 ± 0.017834
Peptide insertion (nm): -1.91186 ± 1.91531
Contacts
Peptide - Water: 70.017500 ± 0.765247
Peptide - Head groups: 0.247500 ± 0.200858
Peptide - Tail groups: 0.0125000 ± 0.0271352
Tilt (°): 91.66250 ± 6.17481
PepDF:
5(ns): CVS
Displacement (nm): 2.0946100 ± 0.0882764
Precession(°): 25.1315 ± 17.6621
50(ns) CVS
Displacement (nm): 6.007820 ± 0.272903
Precession(°): 279.6940 ± 51.3448
100(ns) CVS
Displacement(nm): 8.410650 ± 0.384373
Precession(°): 532.8900 ± 67.4066
200(ns) CVS
Displacement(nm): 11.798000 ± 0.583374
Precession(°): 1002.4800 ± 70.0752
Download JSON File.
5(ns): CVS
Displacement (nm): 2.0946100 ± 0.0882764
Precession(°): 25.1315 ± 17.6621
50(ns) CVS
Displacement (nm): 6.007820 ± 0.272903
Precession(°): 279.6940 ± 51.3448
100(ns) CVS
Displacement(nm): 8.410650 ± 0.384373
Precession(°): 532.8900 ± 67.4066
200(ns) CVS
Displacement(nm): 11.798000 ± 0.583374
Precession(°): 1002.4800 ± 70.0752
Download JSON File.
Peptide Analyses
Peptide Displacement Fingerprint
(PepDF)
Lateral displacement vs
Rotational
Displacement along the trajectory, for different time
windows .
Density maps:
2D-density maps of lipids around the
peptide
along XY and YZ axis, calculated for each lipid type along the
last
microsecond.
Lipid-Peptide Analyses:
z-Position
Z-coordinate, averaged for
differetn
parts of the the system: peptide, membrane, first and
last
backbone (BB) residues and upper of lower leaflet
lipids’
headgroups (HGs).
Minimum distance
Minimum distance (nm) between the
peptide backbone and the lipids (headgroups and
tailgroups).
Number of contacts
Number of contacts between the
peptide backbone and the water or the lipids separated
by
lipid headgroups (HG) or lipid tails, using a cut-off of
0.6
nm.
Lateral density
Lateral density for the different
components of the system: headgroups, tail groups,
peptide
and water.